Manual¶
How to run listed benchmarks¶
After having succesfully installed the basic HPOlib you can download more benchmarks or create your own. Each benchmarks resides in an own directory and consists of an algorithm (+ wrapper if necessary), a configuration file and several hyperparameter configuration descriptions. If you want to use one of the benchmarks listed here, follow these steps:
Let’s say you want to run the Reproducing Kernel Hilbert space (RKHS) function:
RKHS is located with other benchmarks inside
HPOlib/benchmarksfolder. To run the benchmark first go inside that folder.cd HPOlib/benchmarks/rkhs
Inside this folder you can run one the optimizers (smac, tpe or spearmint) on RKHS function using HPOlib :
HPOlib-run -o ../../optimizers/smac/smac_2_10_00-dev -s 23 HPOlib-run -o ../../optimizers/tpe/h -s 23 HPOlib-run -o ../../optimizers/spearmint/spearmint_april2013 -s 23
Or more generally
HPOlib-run /path/to/optimizers/<tpe/hyperopt|smac|spearmint|tpe/random> [-s seed] [-t title]
By default, the optimizers will run 200 evaluations on the function. For smac and tpe this will take about 2 mins but for spearmint it will be longer than 45 mins, so change
number_of_jobsparameter inconfig.cfgfile in same folder to 50 or less.[SMAC] p = params.pcs [TPE] space = space.py [SPEARMINT] config = config.pb [HPOLIB] console_output_delay = 2.0 function = python ../rkhs.py number_of_jobs = 200 #Change this to 50. result_on_terminate = 1000
Now you can plot results for the experiment in different ways:
Plot the results of only one optimizer:
HPOlib-plot FIRSTRUN smac_2_10_00-dev_23_*/smac_*.pkl -s `pwd`/Plots/
The Plots can be found inside folder named
Plotsin current working directory (HPOlib/benchmarks/rkhs)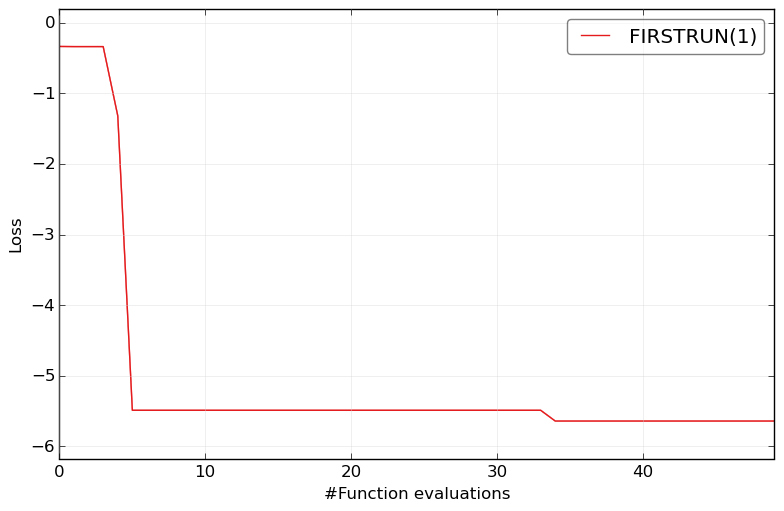
and if you have run all optimizers and want to compare their results:
HPOlib-plot SMAC smac_2_10_00-dev_23_*/smac_*.pkl TPE hyperopt_august2013_mod_23_*/hyp*.pkl SPEARMINT spearmint_april2013_mod_23_*/spear*.pkl -s `pwd`/Plots/
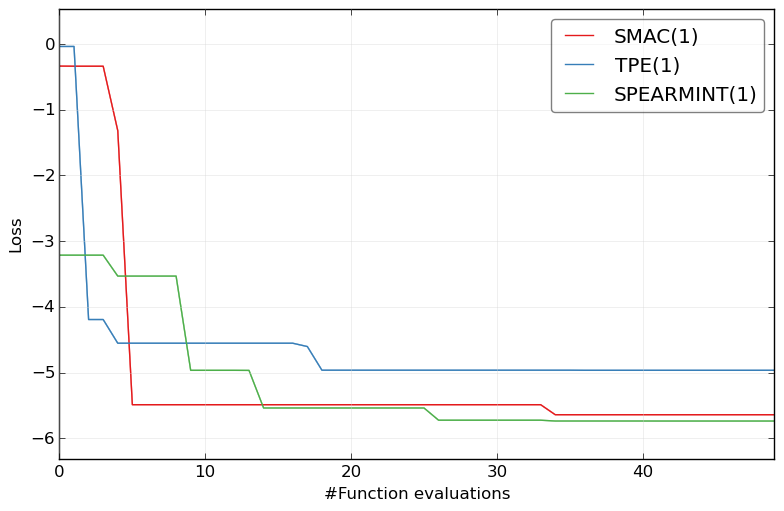
and to check the general performance on this super complex benchmark:
HPOlib-plot RKHS smac_2_10_00-dev_23_*/smac_*.pkl hyperopt_august2013_mod_23_*/hyp*.pkl spearmint_april2013_mod_23_*/spear*.pkl -s `pwd`/Plots/
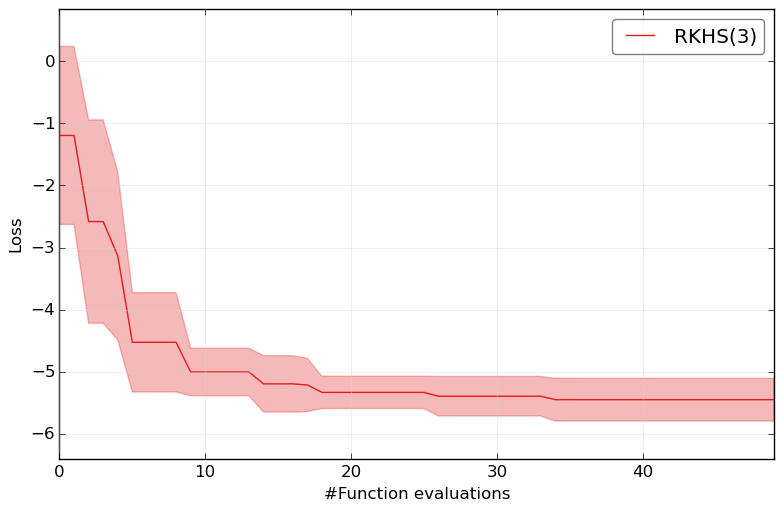
How to run your own benchmarks¶
To run your own benchmark you basically need the software for the benchmark and a search space description for the optimizers smac, spearmint and tpe. In order to work with HPOlib you must put these files into a special directory structure. It is the same directory structure as for the benchmarks which you can download on this website and is explained in the list below. The following lines will guide you through the creation of such a benchmark. Here is a rough guide on what files you need:
- One directory having the name of the optimizer for each optimizer you want to use.
Currently, these are
hyperopt_august2013_mod,random_hyperopt2013_mod,smac_2_10_00-devandspearmint_april2013_mod. - One search space for each optimizer. This must be placed in the directory with the name of the optimizer. You can convert your searchspace to other formats with HPOlib_convert from and to all three different optimizers.
- An executable which implements the HPOlib interface. Alternatively, this can be a wrapper which parser the command line arguments, calls your target algorithm and returns the result to the HPOlib.
- A configuration file config.cfg. See the section on configuring the HPOlib for details.
Example¶
First, create a directory myBenchmark inside the
HPOlib/benchmarks directory. The executable
HPOlib/benchmarks/myBenchmark/myAlgo.py with the target algorithm can
be as easy as
import math
import time
import HPOlib.benchmark_util as benchmark_util
def myAlgo(params, **kwargs):
# Params is a dict that contains the params
# As the values are forwarded as strings you might want to convert and check them
if not params.has_key('x'):
raise ValueError("x is not a valid key in params")
x = float(params["x"])
if x < 0 or x > 3.5:
raise ValueError("x not between 0 and 3.5: %s" % x)
# **kwargs contains further information, like
# for crossvalidation
# kwargs['folds'] is 1 when no cv
# kwargs['fold'] is the current fold. The index is zero-based
# Run your algorithm and receive a result, you want to minimize
result = -math.sin(x)
return result
if __name__ == "__main__":
starttime = time.time()
# Use a library function which parses the command line call
args, params = benchmark_util.parse_cli()
result = myAlgo(params, **args)
duration = time.time() - starttime
print "Result for this algorithm run: %s, %f, 1, %f, %d, %s" % \
("SAT", abs(duration), result, -1, str(__file__))
As you can see, the script parses command line arguments, calls the target function which is implemented in myAlgo, measures the runtime of the target algorithm and prints a return string to the command line. All relevant information is then extracted by the HPOlib. If you write a new algorithm/wrapper script, you must parse the following call:
target_algorithm_executable --fold 0 --folds 1 --params [ [ -param1 value1 ] ]
The return string must take the following form:
Result for this algorithm run: SAT, <duration>, 1, <result>, -1, <additional information>
This return string is not yet optimal and exists for historic reasons. It’s subject to change in one of the next versions of HPOlib.
Next, create HPOlib/benchmarks/myBenchmark/config.cfg,
which is the configuration file. It tells the HPOlib everything about the benchmark and looks like this:
[TPE]
space = mySpace.py
[HPOLIB]
function = python ../myAlgo.py
number_of_jobs = 200
# worst possible result
result_on_terminate = 0
Since the hyperparameter optimization algorithm must know about the variables
and their possible values for your target algorithms, the next step is to
specify these in a so-called search space. Create a new directory
hyperopt_august2013_mod inside the
HPOlib/benchmarks/myBenchmark directory and save
these two lines of python in a file called mySpace.py. If you look at
the config.cfg, we already the use of the newly created search space.
As problems get more complex, you may want to specify more complex search
spaces. It is recommended to do this in the TPE format, then translate it into
the SMAC format which can then be translated into the spearmint format.
More information on how to write search spaces in the TPE format
can be found in this paper and the hyperopt wiki.
from hyperopt import hp
space = {'x': hp.uniform('x', 0, 3.5)}
Now you can run your benchmark with tpe. The command (which has to be
executed from HPOlib/benchmarks/myBenchmark) is
HPOlib-run -o ../../optimizers/tpe/hyperopt_august2013_mod
Further you can run your benchmark with the other optimizers:
mkdir smac
python path/to/hpolib/format_converter/TpeSMAC.py tpe/mySpace.py >> smac/params.pcs
python path/to/wrapping.py smac
mkdir spearmint
python path/to/hpolib/format_converter/SMACSpearmint.py >> spearmint/config.pb
python path/to/wrapping.py spearmint
Configure the HPOlib¶
The config.cfg is a file, which contains necessary settings about your
experiment. It is designed such that as little as possible information needs to be given.
This means all values for optimizers and the wrapping software are set to the default
values, except you want to change them. Default values are stored in a file called
config_parser/generalDefault.cfg. The following table describes the
values you must provide: The file is divided into sections. You only need to
fill in values for the [HPOLIB] section.
| Key | Description |
|---|---|
| function | The executeable for the target algorithm. The path can either be either absolute or relative to an optimizer directory in your benchmark folder (if the executeable is not found you can try to prepend the parent directory to the path) |
| number_of_jobs | number of evaluations that are performed by the
optimizers. NOTE:When using k-fold-crossvalidation,
SMAC will use k * number_of_jobs evaluations |
| result_on_terminate | If your algorithms crashes, is killed, takes too long etc. This result is given to the optimizer. Should be the worst possible, but realistic result for a problem |
An example can be found in the section [adding your own benchmark](manual.html#config_example). The following parameters can be specified:
| Section | Parameter | Default value | Description |
|---|---|---|---|
| HPOLIB | number_cv_folds | 1 |
number of folds for a crossvalidation |
| HPOLIB | max_crash_per_cv | 3 |
If some runs of the crossvalidation fail, stop the crossvalidation for this configuration after max_crash_per_cv failed folds. |
| HPOLIB | remove_target_algorithm_output | True |
Per default, the target algorithm output is deleted. Set to False to keep the output. This is useful for debugging. |
| HPOLIB | console_output_delay | 1.0 |
HPOlib reads the experiment pickle periodically to print the current status to the command line interface. Doing this often can inhibit performance of your hard-drive (espacially if perform a lot of HPOlib experiments in parallel) so you might want to increase this number if you experience delay when accessing your hard drive. |
| HPOLIB | runsolver_time_limit, memory_limit, cpu_limit | Enforce resource limits to a target algorithm run. If these limits are exceeded, the target algorithm will be killed by the runsolver. This can be used to ensure e.g. a runtime per algorithm or make sure an algorithm does not use too much space on a computing cluster. | |
| HPOLIB | total_time_limit | Enforce a total time limit on the hyperparameter optimization. | |
| HPOLIB | leading_runsolver_info | Important when using THEANO and CUDA, see Configure theano for gpu and openBlas usage | |
| HPOLIB | use_HPOlib_time_measurement | True |
When set to True (the default), the runsolver time measurement is saved. Otherwise, the time measured by the target algorithm is saved. |
| HPOLIB | number_of_concurrent_jobs | 1 |
WARNING: this only works for spearmint and SMAC and is not tested! |
| HPOLIB | function_setup | An executable which is called before the first target algorithm call. This can be for example check if everything is installed properly. | |
| HPOLIB | function_teardown | An executable which is called after the last target algorithm call. This can be for example delete temporary directories. | |
| HPOLIB | experiment_directory_prefix | Adds a prefix to the automatically generated experiment directory. Can be useful if one experiments is run several times with different parameter settings. | |
| HPOLIB | handles_cv | This flag determines whether optimization_interceptor or the optimizer handles cross validation. This is only set to 1 for SMAC and must only be used by optimization algorithm developers. |
The following keys change the behaviour of the integrated hyperparameter optimization packages:
| Section | Parameter | Default value Description | |
|---|---|---|---|
| TPE | space | space.py |
Name of the search space for tpe |
| TPE | path_to_optimizer | ./hyperopt_august2013_mod_src |
Please consult the SMAC documentation. |
| SMAC | p | smac/params.pcs |
Please consult the SMAC documentation. |
| SMAC | run_obj | QUALITY |
Please consult the SMAC documentation. |
| SMAC | intra_instance_obj | MEAN |
Please consult the SMAC documentation. |
| SMAC | rf_full_tree_bootstrap | False |
Please consult the SMAC documentation. |
| SMAC | rf_split_min | 10 |
Please consult the SMAC documentation. |
| SMAC | adaptive_capping | false |
Please consult the SMAC documentation. |
| SMAC | max_incumbent_runs | 2000 |
Please consult the SMAC documentation. |
| SMAC | num_iterations | 2147483647 |
Please consult the SMAC documentation. |
| SMAC | deterministic | True |
Please consult the SMAC documentation. |
| SMAC | retry_target_algorithm_run_count | 0 |
Please consult the SMAC documentation. |
| SMAC | intensification_percentage | 0 |
Please consult the SMAC documentation. |
| SMAC | validation | false |
Please consult the SMAC documentation. |
| SMAC | path_to_optimizer | ./smac_2_06_01-dev_src |
Please consult the SMAC documentation. |
| SPEARMINT | config | config.pb |
|
| SPEARMINT | method | GPEIOptChooser |
The spearmint chooser to be used. Please consult the spearmint documentation for possible choices. WARNING: Only the GPEIOptChooser is tested! |
| SPEARMINT | method_args | Pass arguments to the chooser method. Please consult the spearmint documentation for possible choices. | |
| SPEARMINT | grid_size | 20000 |
Length of the Sobol sequence spearmint uses to optimize the Expected Improvement. |
| SPEARMINT | spearmint_polling_time | 3.0 |
Spearmint reads its experiment pickle and checks for finished jobs periodically to find out whether a new job has to be started. For very short functions evaluations, this value can be decreased. Bear in mind that this puts load on your hard drive and can slow down your system if the experiment pickle becomes large (e.g. for the AutoWeka benchmark) or you run a lot of parallel jobs (>100). |
| SPEARMINT | path_to_optimizer | ./spearmint_april2013_mod_src |
The config parameters can also be set via the command line. A use case for this feature is to run the same experiment multiple times, but with different parameters. The syntax is:
HPOlib-run -o spearmint/spearmint_april2013_mod --SECTION:argument value
To set for example the spearmint grid size to 40000, use the following call
HPOlib-run -o spearmint/spearmint_april2013_mod --SPEARMINT:grid_size 40000
If your target algorithm is a python script, you can also load the config file
from within your target algorithm. This allows you to specify extra parameters
for your target algorithm in the config file. Simply import
HPOlib.wrapping_util in your python script and call
HPOlib.wrapping_util.load_experiment_config_file().
The return value is a python config parser object.
Configure theano for gpu and openBlas usage¶
The THEANO-based benchmarks can
be speed-up by either running them on a nvidia GPU or with an optimized BLAS library.
Theano is either configured with theano flags, by changing the value of a variable
in the target program (not recommended as you have to change source code)
or by using a .theanorc file. The .theanorc file is good for
global configurations and you can find more information on how to use it on the
theano config page.
For a more fine-grained control of theano you have to use theano flags.
Unfortunately, setting them in the shell before invoking HPOlib-run
does not work and therefore these parameters have to be added set via the
config variable leading_runsolver_info. This is already set to a
reasonable default for the respective benchmarks but has to be changed in order
to speed up calculations.
For openBlas, change the paths in the following paragraph and replace the value of the
config variable leading_runsolver_info. In case you want to change
more of the theano behaviour (e.g. the compile directory) you must append these
flags to the config variable.
OPENBLAS_NUM_THREADS=2 LD_LIBRARY_PATH=$LD_LIBRARY_PATH:/path/to/the/openBLAS/lib LIBRARY_PATH=$LIBRARY_PATH:/path/to/the/openBLAS/lib THEANO_FLAGS=floatX=float32,device=cpu,blas.ldflags=-lopenblas
If you want to use CUDA on your nvidia GPU, you have to change
device=cpu to device=gpu and add
cuda.root=/usr/local/cuda to the THEANO flags. Change cuda.root
to your cuda installation directory if you did not install cuda to the
default location. For that, replace the path cuda.root=/usr/local/cuda
with the path to your CUDA installation.
How to run your own optimizer¶
Before you integrate your own optimization algorithm, make sure that you know
how the HPOlib is structured and read the section The HPOlib Structure.
The interface to include your own optimizer is straight-forward. Let’s assume
that you have written a hyperparameter optimization package called BayesOpt2.
You tell the HPOlib to use your software with the command line argument
-o or --optimizer. A call to
HPOlib-run -o /path/to/BayesOpt2 should the run
an experiment with your newly written software.
But so far, the HPOlib does not know how to call your software. To let the HPOlib know about the interface to your optimizer, you need to create the three following files (replace BayesOpt2 if your optimization package has a different name):
- BayesOpt2.py: will create all files your optimization package needs in order
- to run
- BayesOpt2_parser.py: a parser which can change the configuration of your
- optimization algorithm based on HPOlib defaults
- BayesOpt2Default.cfg: default configuration for your optimization algorithm
Moreover, your algorithm has to call a script of the HPOlib namely
optimization_interceptor.py, which does bookkeeping and manages a
potential cross validation. The rest of this section will explain how to call
optimization_interceptor.py and the interface your scripts must provide
and the functionality which they must
perform.
Calling optimization_interceptor.py¶
BayesOpt2.py¶
To run BayesOpt2, HPOlib will call the main function of the script
bayesopt2.py. The function signature is as follows:
(call_string, directory) = optimizer_module.main(config=config, options=args, experiment_dir=experiment_dir, experiment_directory_prefix=experiment_directory_prefix)
Argument config is of type ConfigParser,
options of type ArgumentParser
and experiment_dir is a string to the experiment directory. The return
value is a tuple (call_string, directory). call_string must
be a valid (bash) shell command which calls your hyperparameter optimization
package in the way you intend. You can construct the call string based on the
information in the config and the options you are provided with.
directory must be a new directory in which all experiment output will
be stored. HPOlib-run will the change in to the output directory
which your function returned and execute the call string. Your script must
therefore do the following in the main function:
Set up an experiment directory and return the path to the experiment directory. It is highly recommended to create a directory with the following name:
<experiment_directory_prefix><bayesopt2><time_string>
Return a valid bash shell command, which will be used to call your optimizer from the command line interface. The target algorithm you want to optimize is mostly called
optimization_interceptor.py, except for SMAC which handles crossvalidation on its own. Callingoptimization_interceptor.pyallows optimizer independend bookkeeping. The actual function call is the invoked by the HPOlib. Its interface ispython optimization_interceptor.py -param_name1 'param_value' -x '5' -y '3.0'`
etc… The function simply prints the loss to the command line. If your hyperparameter optimization package is written in python, you can also directly call the method
doForTPE(params), where the params argument is a dictionary with all parameter values (both key and value being strings).
Have a look at the bundled scripts smac_2_06_01-dev.py,
spearmint_april2013_mod.py and hyperopt_august2013_mod.py
to get an idea what can/must be done.
BayesOpt2_parser.py¶
The parser file implements a simple interface which only allows the manipulation of the config file:
config = manipulate_config(config)
See the python documentation
for the documentation of the config object. Common usage of
manipulate_config is to check if mandatory arguments are provided.
This is also the recommended place to convert values from the HPOLIB section to
the appropriate values of the optimization package.
BayesOpt2Default.cfg¶
A configuration file for your optimization package as described in the configuration section.
Convert Search Spaces¶

Test/Validate the Best Configuration(s)¶
To get an unbiased performance estimate of the best configuration(s) found, HPOlib offers a script to run a test function with these configurations. The scripts is called like:
HPOlib-testbest --all|--best|--trajectory --cwd path/to/the/optimization/directory
HPOlib-testbest will open the experiment pickle
file which is used for HPOlib bookkeeping, extract the hyperparameters for
the best configuration and call the test function specified in the
configuration file. The result of the test function is then stored in the
experiment pickle and can be further processed. The first argument (either
--all, --best or --trajectory determines for which
configurations the HPOlib will call the test script.
--all: Will call the test-script for all configurations. This is can be very expensive.--best: Call the test-script only for the best configuration.--trajectory: Not yet implemented!
The second argument --cwd tells HPOlib in which experiment directory
it should run test the configurations. As an example, consider the usecase that
we ran SMAC to optimize the logistic regression and want to get
the test performance for the best configuration.
HPOlib-testbest --best --cwd logreg/nocv/smac_2_08_00-master_2000_2014-11-7--16-49-28-166127/
Further options are:
--redo-runs: If argument is given, previous runs will be executed again and the previous results will be overwritten.--n-jobs: Number of parallel function evaluations. You should not set this number higher than the number of cores in your computer.